Locard-Paulet M, Parra J, Albigot R Mouton-Barbosa E, Bardi L, Burlet-Schiltz O, Marcoux J. VisioProt-MS: interactive 2D maps from intact protein mass spectrometry. Bioinformatics. 2018 Aug 6.
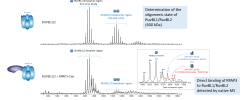
The dynamic inspection of LC-MS 2D maps and the direct comparison of several LC-MS runs with VisioProt-MS facilitates topdown analysis by revealing differences in protein footprints between samples/experimental conditions. This user-friendly application allows the comparison of not only multiple LC-MS runs (including from different deconvolution suites), but also LC-MS and LC-MS/MS runs of the same sample. Its dynamic features enable to pinpoint potential new proteoforms, quickly reject wrongly assigned Protein Spectral Matches (PSM) and spot intense proteoforms that remain unassigned.