Proline
An open source data integration framework and a software suite for mass spectrometry based proteomics:
- Multiple search engines support
- Target/decoy results validation
- Benjamini-Hochberg FDR filtering
- Easy datasets comparison using spectral counting
- Efficient label-free MS quantification
- Isobaric labeling support (TMT, iTraQ)
- PTM sites and clusters exploitation
- User-friendly data visualization
- Highly customizable results exp
ProlineSuite Version 2.3.0
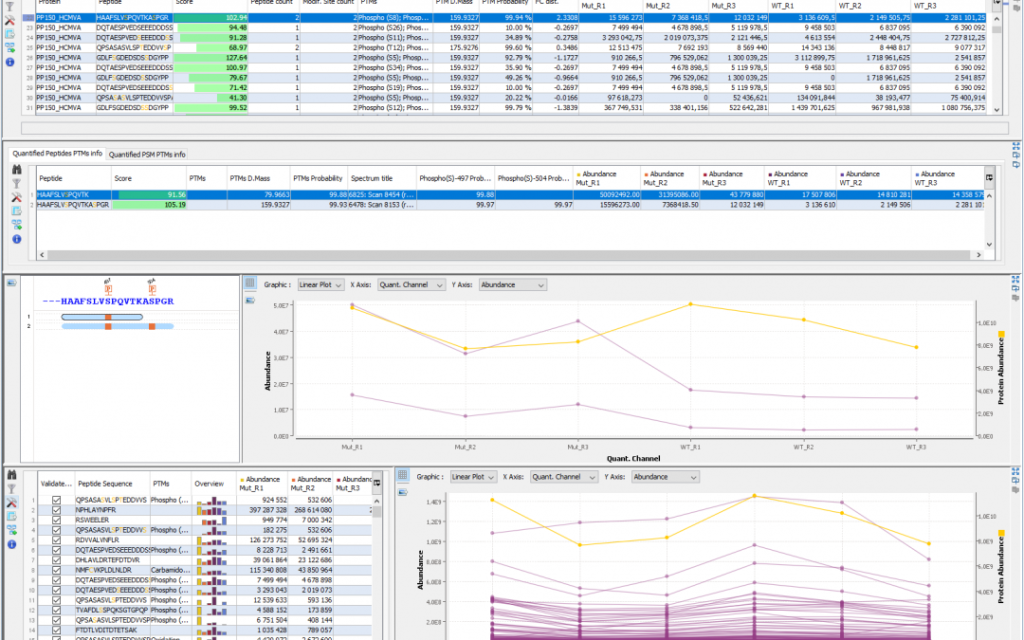
This new release focuses mainly on Isobaric labelling quantification in particular by taking into account purity correction matrices and Precursor Ion Fraction (PIF) information. Works was also carried out on the overall pipeline by developing mzdbConverter, a new tool for converting raw data to mzDB, and MGFBoost to generate more accurate peaklists.
See Release Note for complete description
Proline Version 2.3.3 is available for download here



