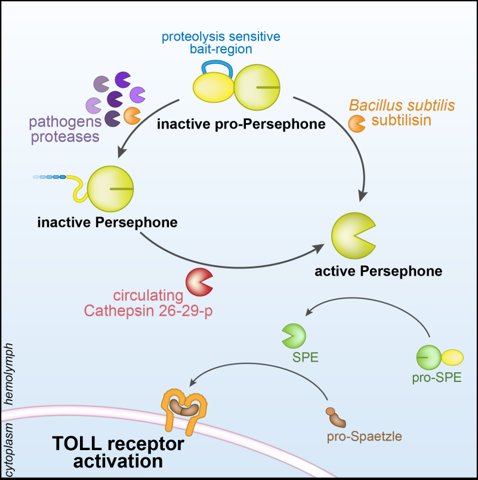
N-terminal oriented proteomics approach
Issa N., Guillaumot N., Lauret E., Matt N., Schaeffer-Reiss C., Van Dorsselaer A., Reichhart J. M., Veillard F., The Circulating
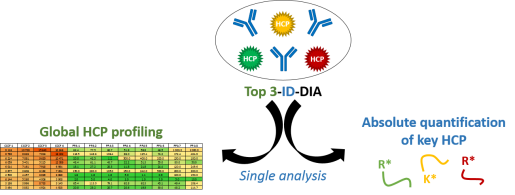
Top 3-ID-DIA method
Husson, G.; Delangle, A.; O’Hara, J.; Cianferani, S.; Gervais, A.; Van Dorsselaer, A.; Bracewell, D.; Carapito, C., Dual Data-Independent Acquisition
cHPP Paper: sperm proteomics
Carapito C, Duek P, Macron C, Seffals M, Rondel K, Delalande F, Lindskog C, Fréour, Vandenbrouck Y, Lane L, Pineau
Proteomics & métabolomics Workshop
ProFI supports the workshop « Analyse et interprétation des données en protéomique et métabolomique : points communs et spécificités ». 28 Mars
Proline Release 1.6 is available !
New validation service (allow to propagate PSM Validation and/or Protein Set filtering), PTM sites identification available for merged dataset, Studio
Thematic course « Quantitative proteomics »
This summer school will take place at Sète, 28 May- 01 June 2018. For more information please contact Myriam Ferro.
Proline Software
If you need some advice or additional training for the use of our software Proline do not hesitate to contact
First meeting FRISBI/ProFI
Mass spectrometry in structural biology November 21, 2017, CNRS, salle La Terrasse, Gif sur Yvette. Click here to find the
First meeting of the French proteomics platforms
Place : Disneyland Paris Date : 2 october 2017
Label-free quantification using Proline
December 2016 : This training session was dedicated to the use of Proline for the analyses of label-free high throughput